Donate processing power to help find a treatment for Covid-19
Help crack Covid
If you want to help stop the spread of Covid-19, wash your hands. Follow medical advice. If you want to help some more, though, it wouldn't hurt to chuck some of your processing power at protein modelling. Folding@home is a program that let's you do exactly that: analysing protein structures to find potential "druggable sites". I won't pretend to fully understand the science, but I do know this is more likely to help than doing nothing.
The program simply runs in the background, using spare CPU to run simulations of coronavirus proteins.
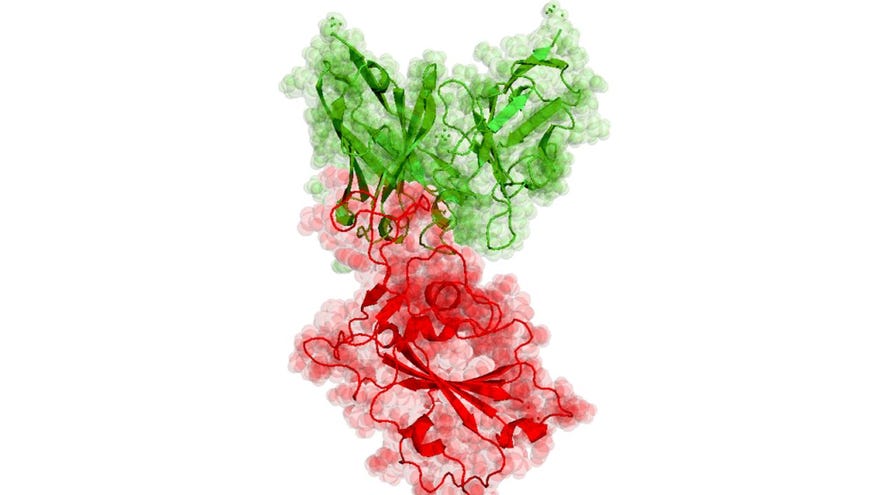
Folding@home Director Greg Bowman, who's also a researcher at Washington University in St. Louis, gives a rough explanation of how it works in this blog post. Viruses use proteins, "molecular machines", to suppress our immune systems and reproduce. Bowman explains that other approaches to modelling these proteins "only reveal a single snapshot of a protein’s usual shape", whereas simulations can give you a moving picture. The goal is to accurately model how the atoms move within the virus's protein structure.
"Doing so can reveal new therapeutic opportunities", Bowman says.
"For example, in our recent paper, we simulated a protein from Ebola virus that is typically considered ‘undruggable’ because the snapshots from experiments don’t have obvious druggable sites. But, our simulations uncovered an alternative structure that does have a druggable site. Importantly, we then performed experiments that confirmed our computational prediction, and are now searching for drugs that bind this newly discovered binding site."
He compares each download to buying a lottery ticket. The more simulations they run, the more likely they are to stumble on something useful. Right now, too many people are chipping in for their servers to handle, so he warns that there will be intermittent downtime.
You can download the software here.
Many gaming events have been cancelled or postponed due to Covid-19, including our dear own Rezzed. Here's a helpful list of every event we know has been affected.